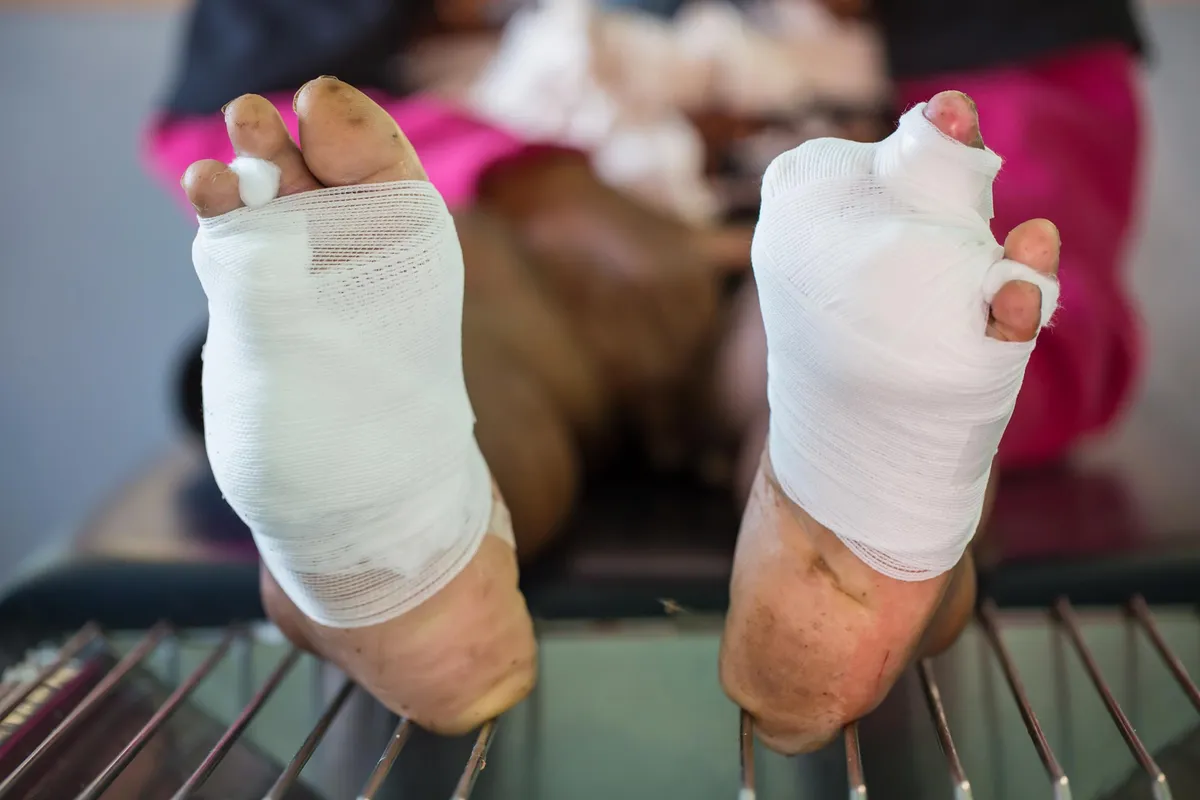
Diabetic foot infections lead to more amputations than most people realize. They're complex, stubborn, and often don't respond well to standard antibiotics. Until now, doctors have known surprisingly little about the specific bacteria driving these infections—especially E. coli, which shows up constantly in wound samples but remains poorly understood.
A team at King's College London decided to look closer. They sequenced the complete DNA of 42 E. coli strains taken from infected diabetic foot ulcers across ten countries: Nigeria, Ghana, the UK, Sweden, Malaysia, China, South Korea, Brazil, India, and the USA. What they found rewrites how we should think about these infections.
The Diversity Problem
The bacteria weren't all the same. The 42 strains belonged to many different genetic groups, each carrying its own combination of resistance genes and virulence factors—the bacterial tools that make infections worse. This matters because it means there's no single culprit to target. Instead, multiple unrelated strains have independently learned to survive in the diabetic foot environment. Some evolve resistance to one antibiotic, others to several. Some are particularly good at attaching to tissue or evading the immune system. Some do both.
We're a new kind of news feed.
Regular news is designed to drain you. We're a non-profit built to restore you. Every story we publish is scored for impact, progress, and hope.
Start Your News DetoxAbout 8 percent of the strains were classified as multidrug-resistant or extensively drug-resistant, meaning they can withstand multiple antibiotics or nearly all available options. That's the kind of finding that keeps infectious disease specialists awake at night.
"Understanding these bacteria at a genomic level is a crucial step towards improving diagnosis and enabling more targeted treatments for people with diabetes," said Vincenzo Torraca, the study's senior author at King's College London. "By identifying which E. coli strains are most common and which antibiotics they are likely to resist, clinicians can choose therapies that are more likely to work."
This precision matters most in places where rapid diagnostic tools aren't available. Low-resource settings see E. coli diabetic foot infections more frequently, yet clinicians often have to guess which antibiotics might work. Knowing which strains are circulating—and what they're resistant to—could mean the difference between saving a limb and losing one.
What Comes Next
The researchers are now studying how specific virulence factors actually function inside diabetic foot wounds. Many of the bacterial samples carried genes for tissue attachment and immune evasion. If scientists can understand how these mechanisms work in the wound environment, they might identify new treatment targets entirely—moving beyond antibiotics toward therapies that disrupt how the bacteria survives.