Scientists have watched a virus do something unexpected: instead of fighting its way into a host cell's existing machinery, it builds a specialized workspace designed just for itself.
Researchers from Kyoto University and collaborators used advanced techniques to track how the giant virus Acanthamoeba polyphaga mimivirus (APMV) replicates inside amoeba cells. What they found challenges how we think about viral infection. The virus doesn't hijack the host's translation system in the straightforward way we'd expect — it creates something closer to a custom workshop.
The Codon Problem
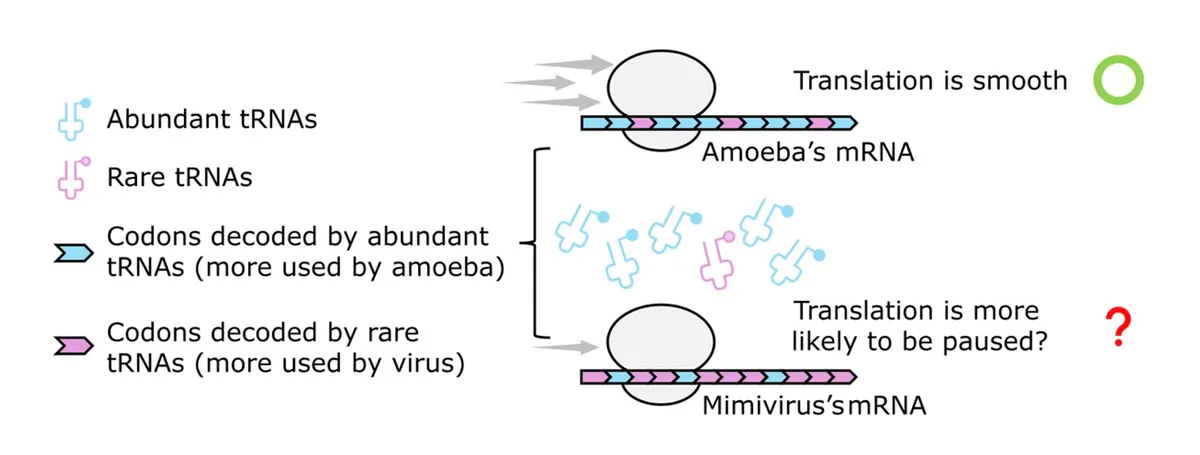
Here's the puzzle the team was trying to solve: viruses need to use the host cell's machinery to translate their genetic instructions into proteins. But many viruses, including this one, use a genetic code that doesn't match their host's. APMV's genome is rich in AT sequences with only 28% GC content, while its amoeba host has 58% GC content. This mismatch should create a bottleneck — like trying to use a specialized tool in a workshop that wasn't designed for it.
We're a new kind of news feed.
Regular news is designed to drain you. We're a non-profit built to restore you. Every story we publish is scored for impact, progress, and hope.
Start Your News DetoxYet when the researchers analyzed ribosome activity using ribosome profiling and tRNA sequencing, they found something surprising. The virus's messenger RNA was translated more smoothly than the host's own mRNA, despite the codon mismatch. The virus wasn't rewriting the host's tRNA pool to favor its own genes. Something else was happening.
The Hidden Workshop
The answer emerged from closer analysis: the virus creates a localized subcellular environment — essentially a specialized organelle — within the amoeba cell. Inside this space, the codons the virus uses frequently are more accessible to matching tRNAs than those same codons are on the host's mRNA. It's like the virus has created a room where its preferred tools are within arm's reach.
This discovery matters beyond amoebas and giant viruses. The researchers suspect this heterogeneous translation strategy — creating a custom molecular environment rather than modifying the whole cell — may be common among many viruses, including human pathogens. It's a fundamentally different approach from how bacterial viruses operate.
"Our study naturally leads to many fascinating questions," says first author Ruixuan Zhang. "How is this subcellular environment created? Which specific proteins or RNAs drive its formation? Can this heterogeneous molecular distribution be generalized to other intracellular microorganisms?"
The team's next step is to map this viral translation environment in more detail, building a systematic picture of how infections actually work at the molecular level. Understanding this mechanism could eventually reshape how we think about treating viral infections — not just stopping viruses, but understanding the precise infrastructure they build to survive.